Votre recherche
Contacter l’IPC
PRENDRE UN RENDEZ-VOUS MÉDICAL
Vous pouvez appeler le Bureau des Rendez-Vous au numéro suivant (SAUF TepScan et Scintigraphie) :
04 91 22 30 30, de 8h à 18h, du lundi au vendredi.
Tous les numéros utiles : cliquez-ici.
Contacter le standard
Pour vos autres demandes (être mis en relation avec un service administratif, le secrétariat d’un médecin pour un compte rendu…), vous pouvez appeler le standard au numéro suivant :
04 91 22 33 33, 24h/24.
Nos actualités
 Semaine européenne de la vaccination 2024 24 avril 2024 - L’Institut Paoli-Calmettes, comme tous les autres centres Unicancer, se mobilise durant cette semaine pour sensibiliser le public et les professionnels de santé à l’importance de la vaccination.
Semaine européenne de la vaccination 2024 24 avril 2024 - L’Institut Paoli-Calmettes, comme tous les autres centres Unicancer, se mobilise durant cette semaine pour sensibiliser le public et les professionnels de santé à l’importance de la vaccination.  Renouvellement de Labellisation – MaRIH – Filière de santé Maladies Rares Immuno-Hématologiques 23 avril 2024 - -- Renouvellement de Labellisation de la filière MaRIH – Filière de santé Maladies Rares Immuno-Hématologiques --
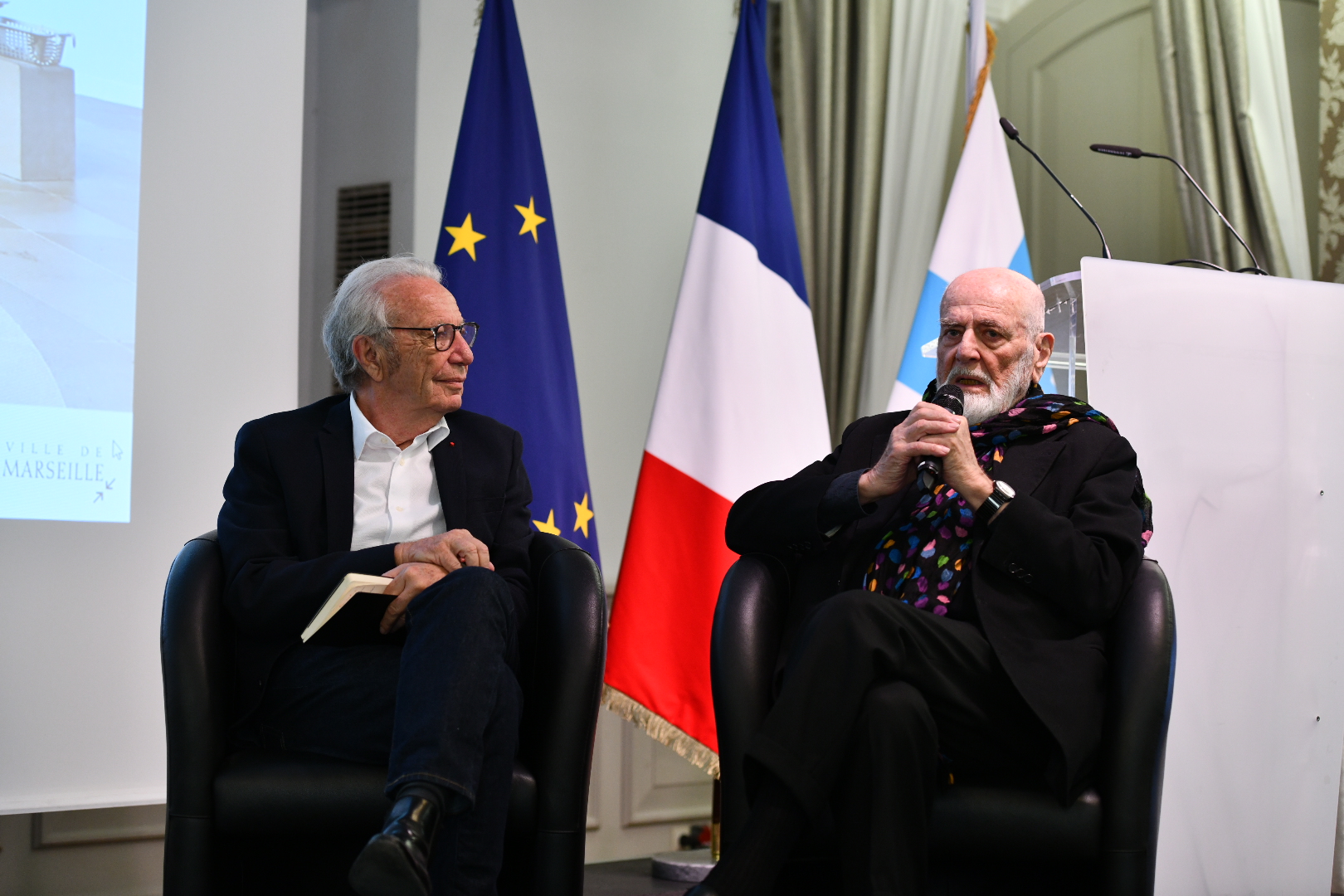
Renouvellement de Labellisation – MaRIH – Filière de santé Maladies Rares Immuno-Hématologiques 23 avril 2024 - -- Renouvellement de Labellisation de la filière MaRIH – Filière de santé Maladies Rares Immuno-Hématologiques --  Retour en images > Art-Santé-Sacré 16 avril 2024 - Découvrez le retour en images de notre Rencontre Art-Santé-Sacré !
Retour en images > Art-Santé-Sacré 16 avril 2024 - Découvrez le retour en images de notre Rencontre Art-Santé-Sacré !